Coronaviruses and their nidovirus relatives, such as arteriviruses, represent a unique and highly relevant group of positive-sense RNA (+RNA) viruses. They possess some of the largest known RNA genomes and employ an exceptionally complex replication strategy that relies on a coordinated interplay between viral enzymes, host cell membranes, and cellular pathways. Over the past two decades, the repeated emergence of highly pathogenic coronaviruses (SARS-CoV, MERS-CoV, and SARS-CoV-2) has underscored their pandemic potential and the urgent need for sustained, fundamental research into their replication and evolution.
Among RNA viruses, nidoviruses are distinguished by their unconventional molecular biology and exceptional evolutionary capacity. Their replication is supported by a repertoire of rare or atypical protein functions, and the intrinsically error-prone nature of their RNA synthesis generates extensive genetic diversity. This feature underpins their rapid adaptation, immune evasion, and capacity for zoonotic transmission. Elucidating these features at the molecular level is therefore critical for the rational design of antiviral therapeutics, vaccines, and broad-spectrum intervention strategies.
Our research currently focuses on four interconnected themes that together help to paint a comprehensive picture of nidovirus replication and its vulnerabilities.
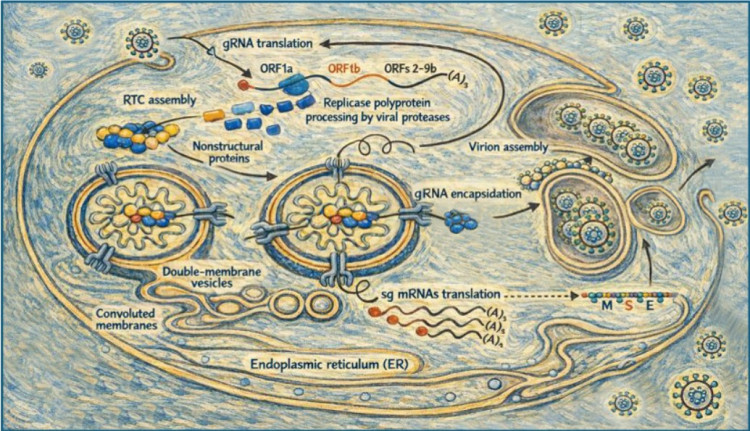
Coronavirus/nidovirus enzymes
 We investigate the structure, function, and regulation of the viral enzymes that drive genome replication, subgenomic mRNA synthesis, and viral RNA processing. These include the RNA-dependent RNA polymerase, viral proofreading and mRNA capping enzymes, and associated enzymatic activities that are uniquely complex among RNA viruses. Detailed biochemical, molecular biological, and structural analyses aim to identify essential functions in coronavirus replication that can be targeted by antiviral compounds.
We investigate the structure, function, and regulation of the viral enzymes that drive genome replication, subgenomic mRNA synthesis, and viral RNA processing. These include the RNA-dependent RNA polymerase, viral proofreading and mRNA capping enzymes, and associated enzymatic activities that are uniquely complex among RNA viruses. Detailed biochemical, molecular biological, and structural analyses aim to identify essential functions in coronavirus replication that can be targeted by antiviral compounds.
Nidovirus replication organelles
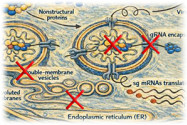
 A defining feature of coronavirus and nidovirus replication is the induction of specialized membrane-bound replication organelles within infected cells. These virus-induced structures are derived from the endoplasmic reticulum and characterized by their double-membrane architecture. They provide a protected environment for viral RNA synthesis, which is connected to the cytosol via a unique molecular pore embedded in te double membrane. Using a combination of biochemistry and advanced electron microscopy, we aim to dissect how these viral mini-organelles are formed, organized, and function during infection.
A defining feature of coronavirus and nidovirus replication is the induction of specialized membrane-bound replication organelles within infected cells. These virus-induced structures are derived from the endoplasmic reticulum and characterized by their double-membrane architecture. They provide a protected environment for viral RNA synthesis, which is connected to the cytosol via a unique molecular pore embedded in te double membrane. Using a combination of biochemistry and advanced electron microscopy, we aim to dissect how these viral mini-organelles are formed, organized, and function during infection.
Lipids and membranes in coronavirus
 Coronaviruses and other nidoviruses hijack host lipid metabolism and intracellular membranes. This feature is closely linked to replication organelle formation, but also to virus entry and assembly. We study how viral transmembrane proteins actively remodel cellular membranes, but also how coronaviruses modulate specific lipid metabolism pathways that appear crucial for efficient replication. This work identifies host-dependent processes that may offer opportunities for host-directed antiviral strategies.
Coronaviruses and other nidoviruses hijack host lipid metabolism and intracellular membranes. This feature is closely linked to replication organelle formation, but also to virus entry and assembly. We study how viral transmembrane proteins actively remodel cellular membranes, but also how coronaviruses modulate specific lipid metabolism pathways that appear crucial for efficient replication. This work identifies host-dependent processes that may offer opportunities for host-directed antiviral strategies.
Antiviral drug development
 Building on fundamental insights into viral replication mechanisms, we develop cell-based assays and experimental models to identify and characterize inhibitors of coronavirus replication. Our work spans target identification, mechanism-of-action studies, and the exploration of antiviral resistance, with a strong emphasis on emerging and high-risk viruses handled under Biosafety Level-3 conditions. A substantial part of this work was and is carried out in the context of EU-funded research programs such a SCORE, CARE, and PANVIPREP.
Building on fundamental insights into viral replication mechanisms, we develop cell-based assays and experimental models to identify and characterize inhibitors of coronavirus replication. Our work spans target identification, mechanism-of-action studies, and the exploration of antiviral resistance, with a strong emphasis on emerging and high-risk viruses handled under Biosafety Level-3 conditions. A substantial part of this work was and is carried out in the context of EU-funded research programs such a SCORE, CARE, and PANVIPREP.
Together, these integrated research themes generate fundamental knowledge that supports antiviral innovation and strengthens preparedness for future coronavirus and nidovirus outbreaks.
Our toolbox includes a broad and complementary set of molecular virological, genetic, biochemical, and structural biology approaches. Genetic platforms to mutate or engineer nidovirus genomes are a cornerstone of our research. They enable the functional analysis of viral genes, proteins, and regulatory elements, facilitate studies of virus–host interactions, and support the generation of tailored recombinant viruses for specific experimental set-ups. Research on highly pathogenic coronaviruses is conducted in the LUMC Biosafety Level-3 (BSL-3) facility, which is managed and operated by members of our team.



